The Pangenome: A Statistical Model
The pangenome, representing the complete genetic repertoire of a species, has become a central concept in modern prokaryotic genomics, moving the field beyond the limitations of a single reference genome. This review provides a comprehensive overview of the pangenomic landscape, from its conceptual foundations to its diverse applications. We critically examine the methodological and statistical underpinnings of pangenome analysis, emphasizing that the pangenome is not a fixed biological entity but a statistical model whose outputs are fundamentally dependent on dataset quality, curation, and taxonomic resolution. We discuss the transition from the simple open/closed dichotomy to more nuanced, ecologically-driven models like pangenome fluidity, which better explain the evolutionary dynamics of prokaryotic genomes. With a special focus on the archaeal pangenome, we highlight the unique challenges and novel evolutionary insights emerging from this under-explored domain. As the field moves towards graph-based representations, AI-driven analysis, and community-level metapangenomics, a rigorous and critical approach to data interpretation will be paramount to unlocking a true understanding of microbial evolution and function.
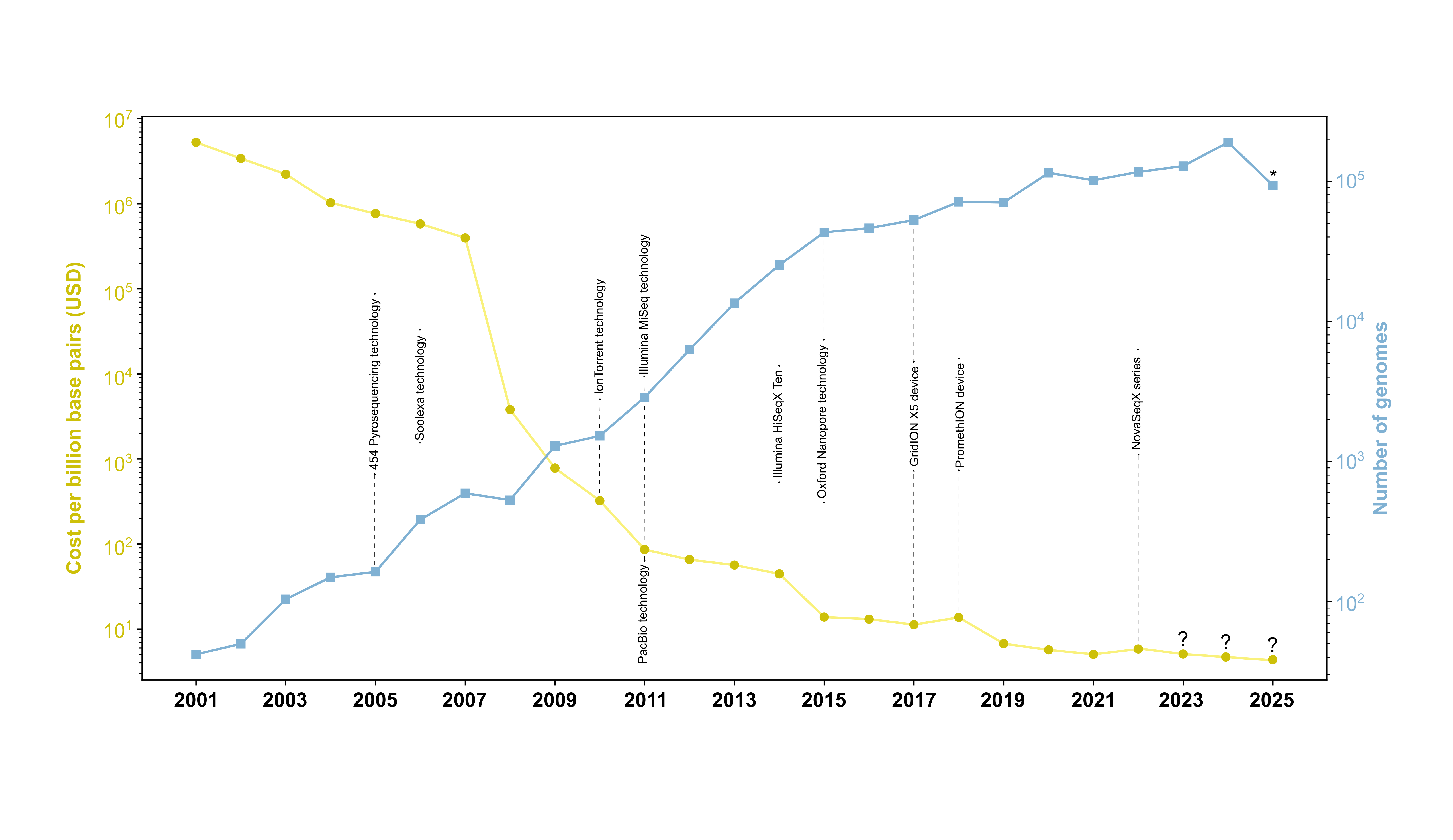
Figure 1. Schematic representation comparing the cost per billion base pairs (USD) and the number of genomes submitted in the NCBI database from 2001 to 2025. The cost trend is depicted in yellow, while the cumulative number of genomes is shown in blue. Both axes use logarithmic scales to facilitate data comparison. The "?" symbol indicates projected data or ideal trends where current data are unavailable, while "*" denotes that the values for 2025 are incomplete, but the expected trend is an ongoing increase. Key milestones in sequencing technology development are highlighted at specific years.
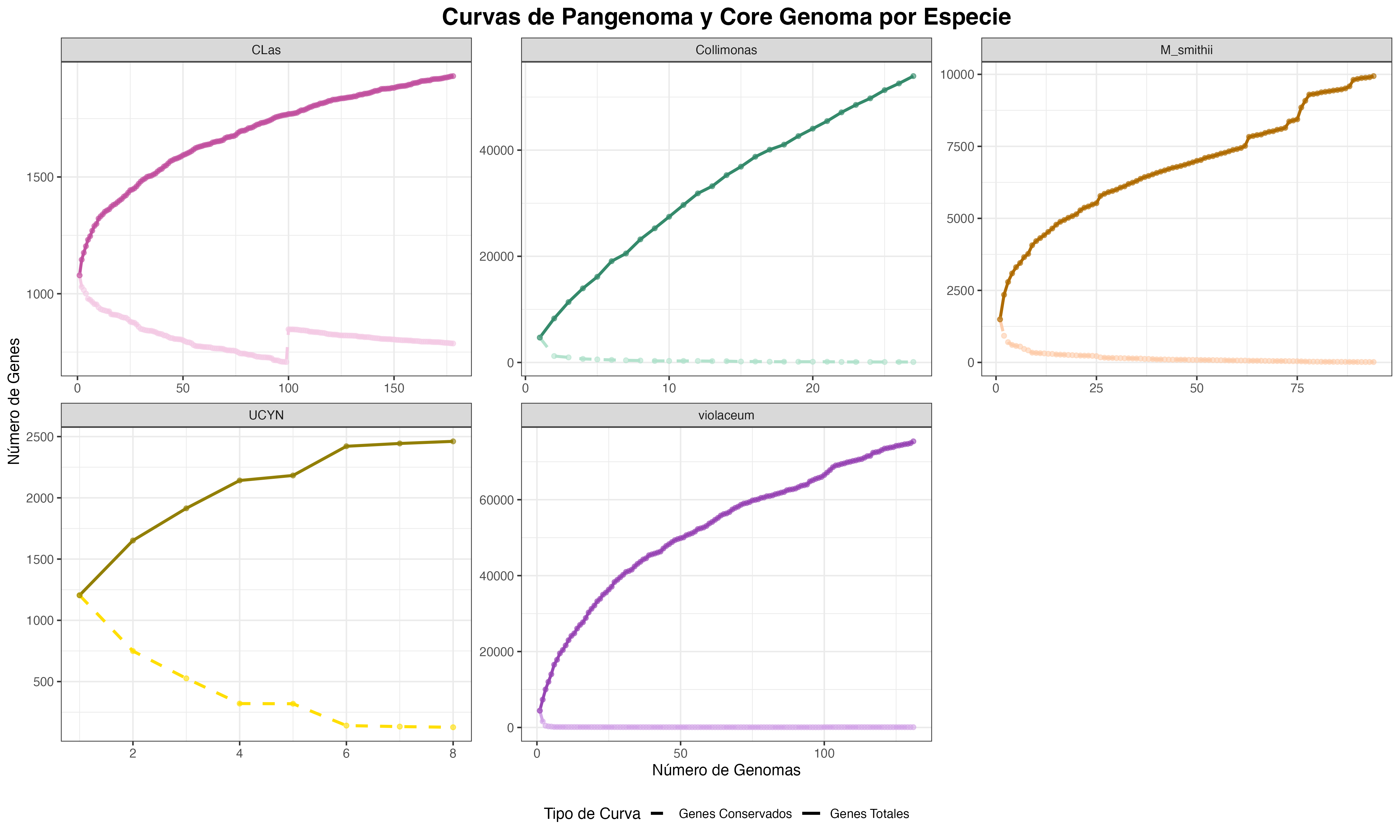
Fig. 2: Comparative pangenome analysis of five prokaryotic taxa with different lifestyles. The figure displays the pangenome growth curve (total genes, solid line) and the core genome decay curve (conserved genes, dashed line) for the genera Chromobacterium, Collimonas, Methanobrevibacter (M. smithii and M. intestinalis), and Candidatus Atelocyanobacterium, and for the species Candidatus Liberibacter asiaticus. In each panel, the x-axis represents the number of genomes added sequentially, and the y-axis represents the number of genes. The openness of each pangenome was modeled using Heaps' law (P=kNλ). Values of λ<0.3 suggest a closed pangenome, while values of λ>0.3 indicate an intermediate open pangenome.
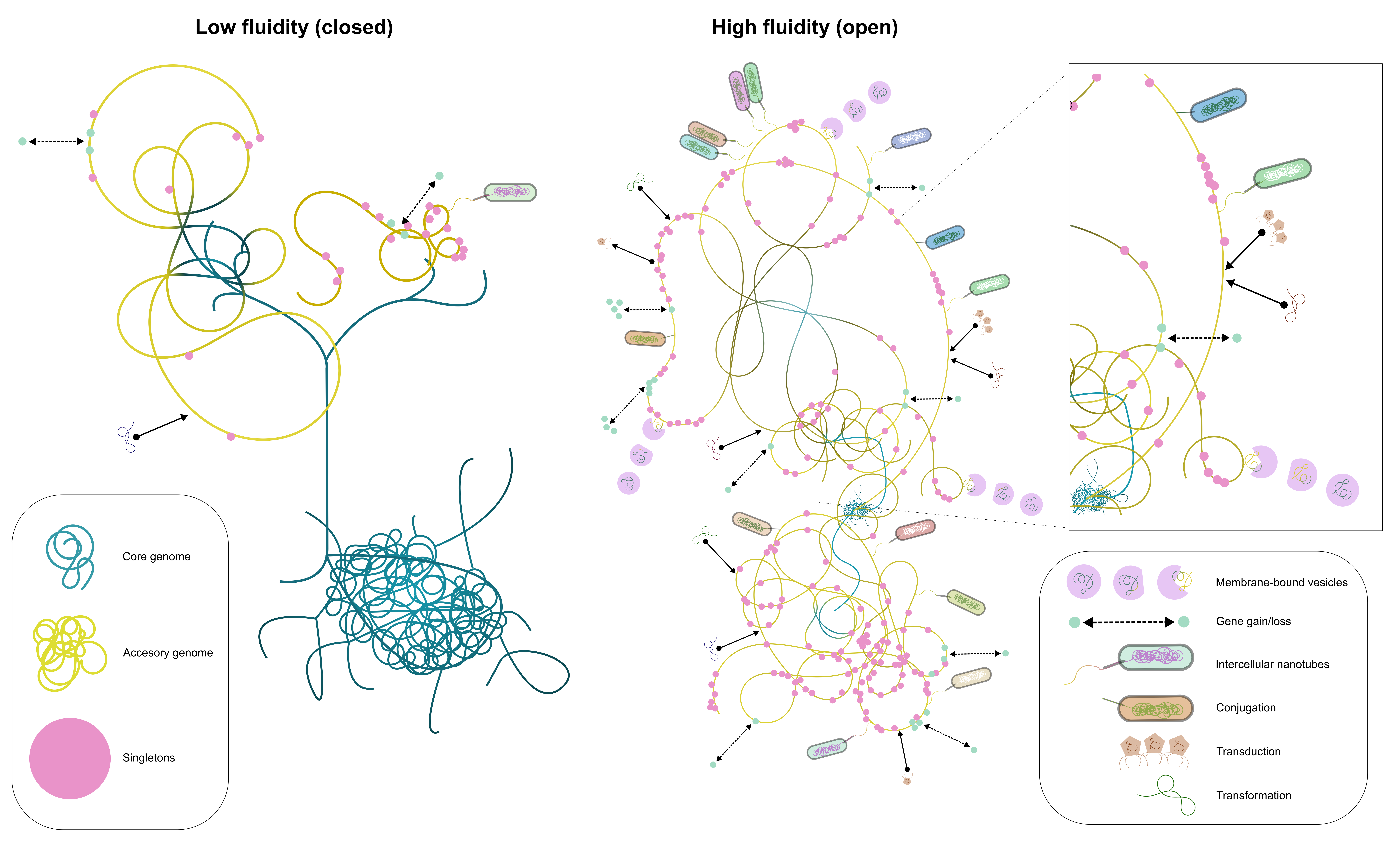
Fig. 3: The pangenome fluidity spectrum driven by horizontal gene transfer (HGT). (A) Low-fluidity (closed) pangenome characterized by a large, stable core genome and a limited accessory genome. (B) High-fluidity (open) pangenome displaying a reduced core genome, an expansive accessory genome, and numerous singletons. The core genome is shown in blue, the accessory genome in yellow, and singletons in pink. The inset box (top right) provides a magnified view of the diverse HGT mechanisms that drive pangenome fluidity, including transformation, transduction, conjugation, membrane-bound vesicles, and intercellular nanotubes.